JANUARY 19th 2023
It’s been quite a long while between blog posts – 3 years that have included a pandemic and many changes to the way we work. I won’t try to cover everything that has happened in the lab in that time, but thought it might be worth summarising some of the highlights.
2022
It felt like some of our longer-term studies were finalised last year and we were joined by new students. We were thrilled to be joined by Pranita and Thomas that will work on regulatory RNA and RNA-binding proteins in EHEC for the PhDs. Their enthusiasm has already made a significant impact on our research.
Peri also joined the lab for her honours year, and received first class for her thesis looking at the targeting specficity of Cas7-11. The project encountered some unexpected set-backs but her hard work and dedication saw the project through, and we wish her all the best in her future endeavors.
Conference travel was back on the table in 2022 and the lab attended the ASM conference in 2022 at the ICC in Sydney, where Pranita won a poster prize. We also travelled to Brisbane for BacPath which was a highlight of the year and Winton received an oral presentation award. We are proud of all our members for their great achievements and for representing the lab in a professional manner.
This year Daniel published his work on the sRNA interactome of MRSA in Nature Communications, this has been a major focus of the lab for a number of years and it’s wonderful to have work published. This work was a collaboration with the Stinear lab at UMelb and the Granneman lab at the Uni of Edinburgh, UK. There is a link to both Nature Comms papers on the Publications page.
Winton submitted his PhD thesis on the sRNA interactome of vancomycin tolerant Staphylococcus aureus. Congratulations! Winton will start as a post-doc in the lab in 2023 working on a joint project on Streptococcus pyogenes.
We are also happy to announce that the lab has been funded to support RNA biology work in Streptococcus pyogenes with Martina Sanderson-Smith (Uni of Wollongong), Mark Davies (Uni Melbourne), and the Walker lab at UQ on an NHMRC Ideas grant. This grant will allow us to expand our research and understand how SNPs in a non-coding RNA drive increased toxin production.

2021
One of the highlights of the year was the ASM conference in 2021, which was held virtually due to the ongoing pandemic. Despite the challenges posed by the virtual format, the lab was able to participate in the conference. This was true for a number of other virtual conferences including the Regulating with RNA in Bacteria and Archaea conference, and ASM World Microbe Forum. Hats off to the conference organisers – it really felt like a brave new world and they worked surprisingly well. That said, it was great to finally see everyone in-person in 2022.
In addition to the conference, the lab was also fortunate to welcome Ken, Serena, and Sam as honours students. These talented students made significant contributions to the lab’s research and all graduated with first class honours. We are proud of their hard work and dedication, and wish them all the best in their future endeavors.
One of the most exciting developments of the year was the lab being awarded an ARC Discovery grant. This grant will allow us to use machine learning to understand sequences and structures that drive sRNA function in E. coli. This work is a collaboration with A/Prof Fatemeh Vafaee at UNSW and we’re really looking forward to seeing how we can dissect our CRAC and CLASH data using new approaches being employed in Fatemeh’s lab.

2020
Good grief, what a year this was.
In early Feb we had the first indicates that 2020 might be a bit different – the borders to Australia were closed around mid-March and all teaching and research had to be completed remotely. Over the next 2 years the borders stayed closed and we were in and out of lockdowns with travel restrictions.
Despite the difficulties of online teaching and research, the year had a number of silver linings. Brandon’s PhD work was published in PNAS and he started a post-doctoral position at the Garvan Institute working on the RNA biology of neurons.
Daniel Neville joined the lab in Feb for his Honours year, and due the lockdowns -had to pivot his project to incorporate a significant amount of EHEC RNA sequencing data analysis into his thesis, highlighting his ability to adapt to unexpected obstacles and continue to produce high-quality work. Daniel identified a number of new 3′ UTRs sRNAs using our Term-seq and Hfq and RNase E CRAC/CLASH data and we hope to be able to complete this work in the near future. He received first class for his work over this challenging year.

SEPTEMBER 8, 2019
Congratulations Brandon on submitting your PhD thesis!
2019 has been and eventful year and last week we celebrated Brandon’s thesis submission. While this is only the beginning of Brandon’s thesis examination, it marks the completion of a large body of work that he should be very proud of.
We’ve had 2 new arrivals this year. In January, Dr Daniel Mediati joined us from UTS. Daniel will be studying RNA-based gene regulation in MRSA. Earlier this year Sylvania also joined the lab to start her PhD looking at gene regulation in MRSA. Welcome (back) to the lab Daniel and Sylvania! We’re all excited about the discoveries you’ll make over the next few years.
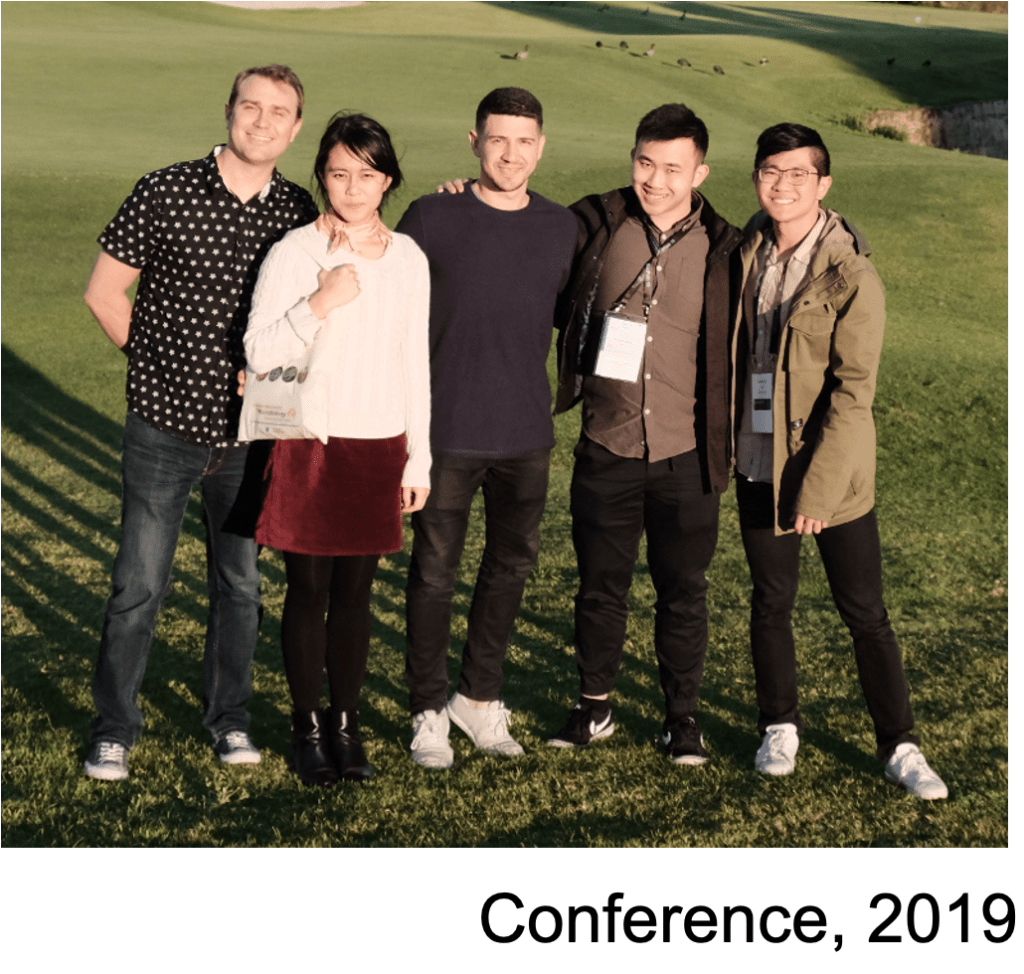
It’s conference season again for the Tree Lab. We’ll all be attending BacPath15 in Perth this year. Daniel, Brandon, and I will be speaking about RNA-based gene regulation in Shiga toxigenic E. coli and MRSA. Sylvania and Winton will be presenting posters on their work in MRSA. The conference is always lots of fun and this year looks to be no exception. Shortly after BacPath15, I will be attending the EMBL non-coding genome conference and visiting a few bacterial RNA groups in Europe. It will terrific opportunity to catch up on the latest and greatest in RNA biology.
Finally, the lab hosted A/Prof Kate Seib from the Institute for Glycomics at Grffith University last week. Kate spoke about her work on developing a gonococcal vaccine and prospects for eradicating this STI. Her talk was extremely well received and we’re all looking forward to reading more about her progress in this area over the coming years.

DECEMBER 7, 2018
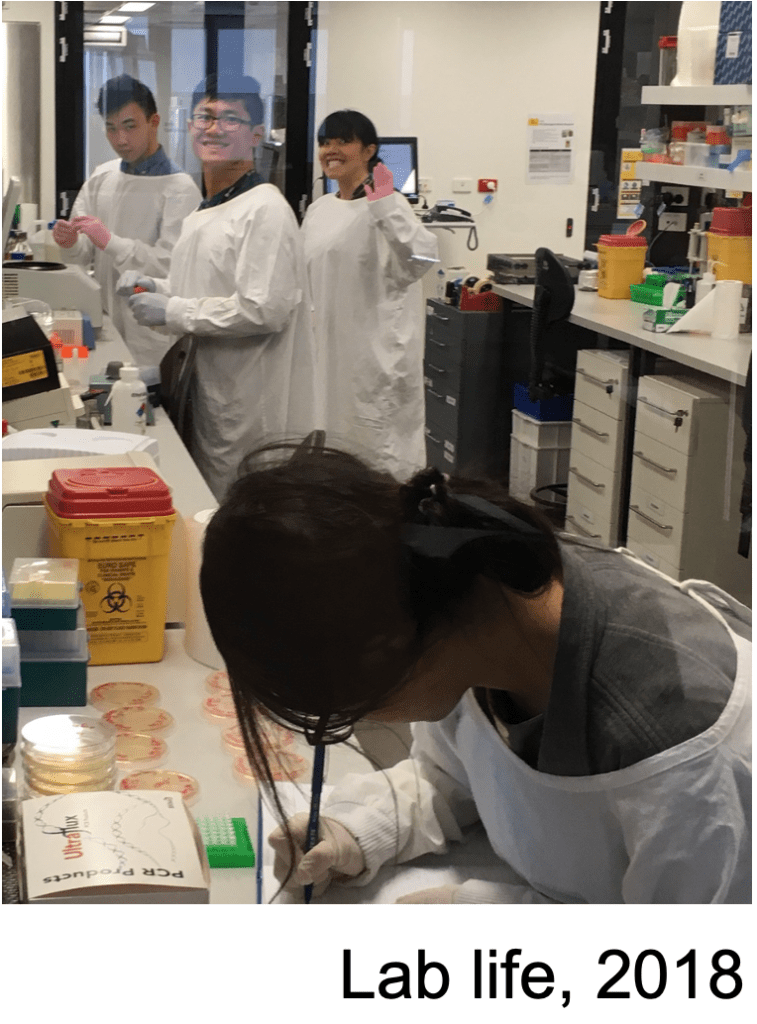
As 2018 draws to a close …
Another successful year! Congratulations to Sylvania Wu on her first class Honours, and welcome back to the lab Winton Wu after a brief sojourn at the Protein Expression Facility @ the University of Queensland. Winton started his PhD in August and is already off to a strong start.
The lab was also very pleased to hear that funding for our EHEC work had been renewed by the NHMRC. These funds will allow us to expand out network analysis of EHEC regulatory RNAs (published last year in EMBO J), and provide a detailed analysis of some of the more significant regulatory interactions that we have found in these networks.

DECEMBER 7, 2017
NHMRC funding 2018-2020
We were very pleased to find out that our application to the NHMRC for Project funding was successful this year. The funds allow us to explore small RNA regulated antibiotic tolerance in multi-drug resistant Staphylococcus aureus.
SEPTEMBER 14, 2017
Spring is springing
The warm weather is back in Sydney (!!). The LaboraTree headed out for a BBQ in Coogee – followed by some Giant Jenga at the Coogee Pavilion.
Brandon and Julia will be giving talks on their work on small RNA function in EHEC and MRSA at BacPath14! Congrats guys! We’re all really excited about his conference – it’s always fun and informative.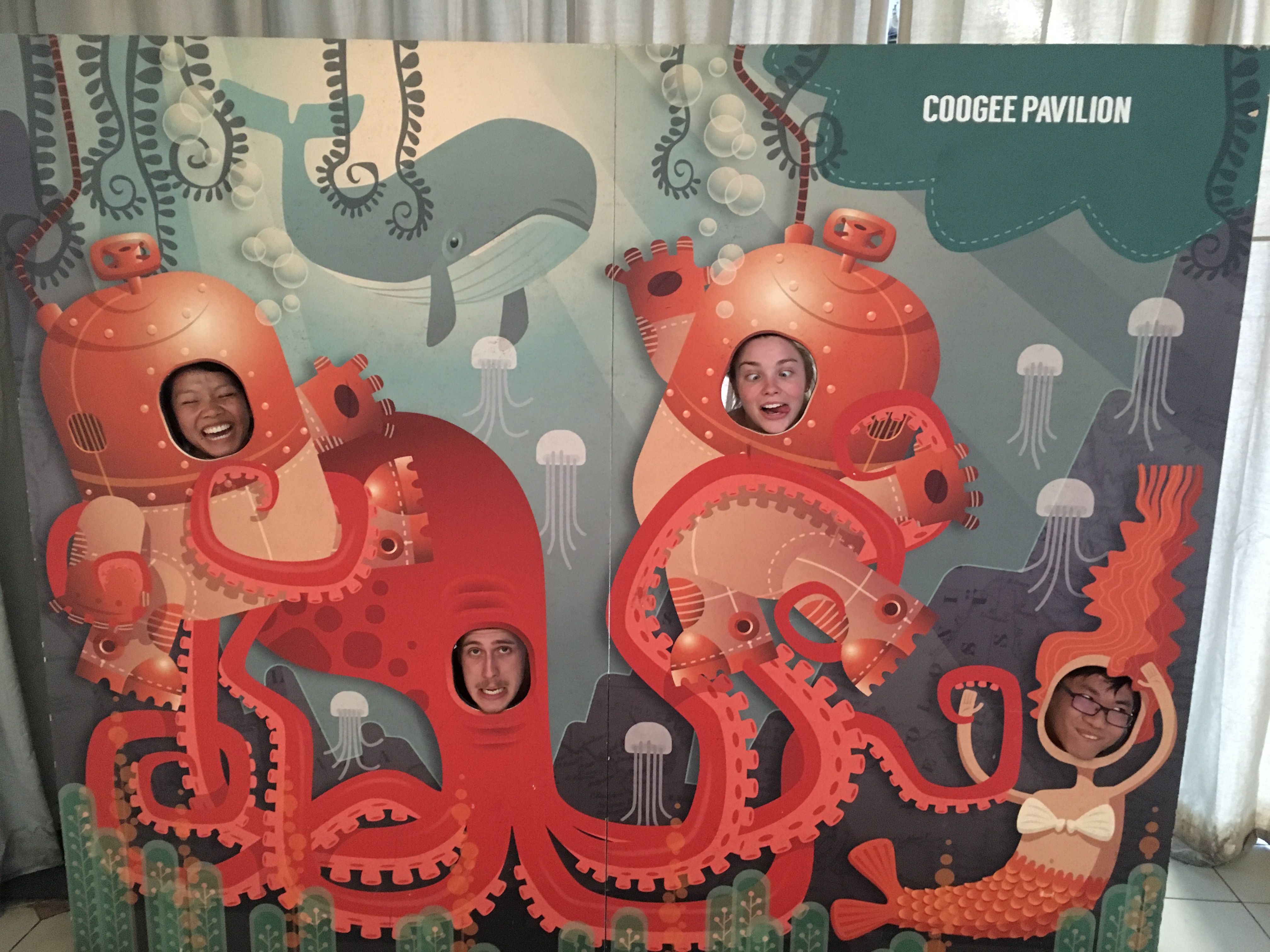
FEBRUARY 19, 2017
Honours 2017
A belated congratulations to Winton Wu on his first class Honours in 2016! Well Done. We’re very excited to welcome Lawrence and Rebecca to the LaboraTree in March 2017 to start their Honours research. Exciting discoveries ahead!